In the Jeopardy! game show contestants are presented with questions formulated as answers that require answers in the form questions. For example, if a contestant selects “Normality for $200” she might be shown the following clue:
“The average ,”
to which she would reply “What is the maximum likelihood estimate for the mean of n independent identically distributed Gaussian random variables from which samples have been obtained?” Host Alex Trebek would immediately exclaim “That is the correct answer for $200!”
The process of doing mathematics involves repeatedly playing Jeopardy! with oneself in an unending quest to understand everything just a little bit better. The purpose of this blog post is to provide an exposition of how this works for understanding principal component analysis (PCA): I present four Jeopardy clues in the “Normality” category that all share the same answer: “What is principal component analysis?” The post was motivated by a conversation I recently had with a well-known population geneticist at a conference I was attending. I mentioned to him that I would be saying something about PCA in my talk, and that he might find what I have to say interesting because I knew he had used the method in many of his papers. Without hesitation he replied that he was well aware that PCA was not a statistical method and merely a heuristic visualization tool.
The problem, of course, is that PCA does have a statistical interpretation and is not at all an ad-hoc heuristic. Unfortunately, the previously mentioned population geneticist is not alone; there is a lot of confusion about what PCA is really about. For example, in one textbook it is stated that “PCA is not a statistical method to infer parameters or test hypotheses. Instead, it provides a method to reduce a complex dataset to lower dimension to reveal sometimes hidden, simplified structure that often underlie it.” In another one finds out that “PCA is a statistical method routinely used to analyze interrelationships among large numbers of objects.” In a highly cited review on gene expression analysis PCA is described as “more useful as a visualization technique than as an analytical method” but then in a paper by Markus Ringnér titled the same as this post, i.e. What is principal component analysis? in Nature Biotechnology, 2008, the author writes that “Principal component analysis (PCA) is a mathematical algorithm that reduces the dimensionality of the data while retaining most of the variation in the data set” (the author then avoids going into the details because “understanding the details underlying PCA requires knowledge of linear algebra”). All of these statements are both correct and incorrect and confusing. A major issue is that the description by Ringnér of PCA in terms of the procedure for computing it (singular value decomposition) is common and unfortunately does not shed light on when it should be used. But knowing when to use a method is far more important than knowing how to do it.
I therefore offer four Jeopardy! clues for principal component analysis that I think help to understand both when and how to use the method:
1. An affine subspace closest to a set of points.
Suppose we are given n numbers as in the initial example above. We are interested in finding the “closest” number to these numbers. By “closest” we mean in the sense of total squared difference. That is, we are looking for a number
such that
is minimized.
This is a (straightforward) calculus problem, solved by taking the derivative of the function above and setting it equal to zero. If we let then
and setting
we can solve for
to obtain
.
The right hand side of the equation is just the average of the n numbers and the optimization problem provides an interpretation of it as the number minimizing the total squared difference with the given numbers (note that one can replace squared difference by absolute value, i.e. minimization of , in which case the solution for m is the median; we return to this point and its implications for PCA later).
Suppose that instead of n numbers, one is given n points in . That is, point i is
. We can now ask for a point
with the property that the squared distance of
to the n points is minimized. This is asking for
.
The solution for can be obtained by minimizing each coordinate independently, thereby reducing the problem to the simpler version of numbers above, and it follows that
.
This is 0-dimensional PCA, i.e., PCA of a set of points onto a single point, and it is the centroid of the points. The generalization of this concept provides a definition for PCA:
Definition: Given n points in , principal components analysis consists of choosing a dimension
and then finding the affine space of dimension k with the property that the squared distance of the points to their orthogonal projection onto the space is minimized.
This definition can be thought of as a generalization of the centroid (or average) of the points. To understand this generalization, it is useful to think of the simplest case that is not 0-dimensional PCA, namely 1-dimensional PCA of a set of points in two dimensions:
 In this case the 1-dimensional PCA subspace can be thought of as the line that best represents the average of the points. The blue points are the orthogonal projections of the points onto the “average line” (see, e.g., the red point projected orthogonally), which minimizes the squared lengths of the dashed lines. In higher dimensions line is replaced by affine subspace, and the orthogonal projections are to points on that subspace. There are a few properties of the PCA affine subspaces that are worth noting:
In this case the 1-dimensional PCA subspace can be thought of as the line that best represents the average of the points. The blue points are the orthogonal projections of the points onto the “average line” (see, e.g., the red point projected orthogonally), which minimizes the squared lengths of the dashed lines. In higher dimensions line is replaced by affine subspace, and the orthogonal projections are to points on that subspace. There are a few properties of the PCA affine subspaces that are worth noting:
- The set of PCA subspaces (translated to the origin) form a flag. This means that the PCA subspace of dimension k is contained in the PCA subspace of dimension k+1. For example, all PCA subspaces contain the centroid of the points (in the figure above the centroid is the green point). This follows from the fact that the PCA subspaces can be incrementally constructed by building a basis from eigenvectors of a single matrix, a point we will return to later.
- The PCA subspaces are not scale invariant. For example, if the points are scaled by multiplying one of the coordinates by a constant, then the PCA subspaces change. This is obvious because the centroid of the points will change. For this reason, when PCA is applied to data obtained from heterogeneous measurements, the units matter. One can form a “common” set of units by scaling the values in each coordinate to have the same variance.
- If the data points are represented in matrix form as an
matrix
, and the points orthogonally projected onto the PCA subspace of dimension k are represented as in the ambient p dimensional space by a matrix
, then
. That is,
is the matrix of rank k with the property that the Frobenius norm
is minimized. This is just a rephrasing in linear algebra of the definition of PCA given above.
At this point it is useful to mention some terminology confusion associated with PCA. Unfortunately there is no standard for describing the various parts of an analysis. What I have called the “PCA subspaces” are also sometimes called “principal axes”. The orthogonal vectors forming the flag mentioned above are called “weight vectors”, or “loadings”. Sometimes they are called “principal components”, although that term is sometimes used to refer to points projected onto a principal axis. In this post I stick to “PCA subspaces” and “PCA points” to avoid confusion.
Returning to Jeopardy!, we have “Normality for $400” with the answer “An affine subspace closest to a set of points” and the question “What is PCA?”. One question at this point is why the Jeopardy! question just asked is in the category “Normality”. After all, the normal distribution does not seem to be related to the optimization problem just discussed. The connection is as follows:
2. A generalization of linear regression in which the Gaussian noise is isotropic.
PCA has an interpretation as the maximum likelihood parameter of a linear Gaussian model, a point that is crucial in understanding the scope of its application. To explain this point of view, we begin by elaborating on the opening Jeopardy! question about Normality for $200:
The point of the question was that the average of n numbers can be interpreted as a maximum likelihood estimation of the mean of a Gaussian. The Gaussian distribution is
. Given the numbers
, the likelihood function is therefore
. The maximum of this function is the same as the maximum of its logarithm, which is
. Therefore the problem of finding the maximum likelihood estimate for the mean is equivalent to that of finding the minimum of the function
. This is exactly the optimization problem solved by 0-dimensional PCA, as we saw above. With this calculation at hand, we turn to the statistical interpretation of least squares:
Given n points in the plane (see figure above), the least squares line
(purple in figure) is the one that minimizes the sum of the squares
. That is, the least squares line is the one minimizing the sum of the squared vertical distances to the points. As with the average of numbers, the least squares line has a statistical interpretation: Suppose that there is some line
(black line in figure) that is unknown, but that “generated” the observed points, in the sense that each observed point i was obtained by perturbing the point
vertically by a random amount from a single Gaussian distribution with mean 0 and variance
. In the figure, an example is shown where the blue point on the unknown line “generates” the observed red point; the Gaussian is indicated with the blue streak around the point. Note that the model specified so far is not fully generative, as it depends on the hidden points
and there is no procedure given to generate the
. This can be done by positing that the
are generated from a Gaussian distribution along the line
(followed by the points
generated by Gaussian perturbation of the y coordinate on the line). The coordinates
can then be deduced directly from the observed points as the Gaussian perturbations are all vertical. The relationship between the statistical model just described and least squares is made precise by a theorem (which we state informally, but is a special case of the Gauss-Markov theorem):
Theorem (Gauss-Markov): The maximum likelihood estimate for the line (the parameters m and b) in the model described above correspond to the least squares line.
The proof is analogous to the argument given for the average of numbers above so we omit it. It can be generalized to higher dimensions where it forms the basis of what is known as linear regression. In regression, the are known as independent variables and
the dependent variable. The generative model provides an interpretation of the independent variables as fixed measured quantities, whereas the dependent variable is a linear combination of the independent variables with added noise. It is important to note that the origins of linear regression are in physics, specifically in work of Legendre (1805) and Gauss (1809) who applied least squares to the astronomical problem of calculating the orbits of comets around the sun. In their application, the independent variables were time (for which accurate measurements were possible with clocks; by 1800 clocks were accurate to less than 0.15 seconds per day) and the (noisy) dependent variable the measurement of location. Linear regression has become one of the most (if not the most) widely used statistical tools but as we now explain, PCA (and its generalization factor analysis), with a statistical interpretation that includes noise in the
variables, seems better suited for biological data.
The statistical interpretation of least squares can be extended to a similar framework for PCA. Recall that we first considered a statistical interpretation for least squares where an unknown line “generated” the observed points, in the sense that each observed point i was obtained by perturbing the point
vertically by a random amount from a single Gaussian distribution with mean 0 and variance
. PCA can be understood analogously by replacing vertically by orthogonally (this is the probabilistic model of Collins et al., NIPS 2001 for PCA). However this approach is not completely satisfactory as the orthogonality of the perturbation is is not readily interpretable. Stated differently, it is not obvious what physical processes would generate points orthogonal to a linear affine subspace by perturbations that are always orthogonal to the subspace. In the case of least squares, the “vertical” perturbation corresponds to noise in one measurement (represented by one coordinate). The problem is in naturally interpreting orthogonal perturbations in terms of a noise model for measurements. This difficulty is resolved by a model called probabilistic PCA (pPCA), first proposed by Tipping and Bishop in a Tech Report in 1997, and published in the J. of the Royal Statistical Society B 2002, and independently by Sam Roweis, NIPS 1998, that is illustrated visually in the figure below, and that we now explain:
In the pPCA model there is an (unknown) line (affine space in higher dimension) on which (hidden) points (blue) are generated at random according to a Gaussian distribution (represented by gray streak in the figure above, where the mean of the Gaussian is the green point). Observed points (red) are then generated from the hidden points by addition of isotropic Gaussian noise (blue smear), meaning that the Gaussian has a diagonal covariance matrix with equal entries. Formally, in the notation of Tipping and Bishop, this is a linear Gaussian model described as follows:
Observed random variables t are given by where x are latent (hidden) random variables, W is a matrix describing a subspace and
are the latent points on an affine subspace (
corresponds to a translation). Finally,
is an error term, given by a Gaussian random variable with mean 0 and covariance matrix
. The parameters of the model are
and
. Equivalently, the observed random variables are themselves Gaussian, described by the distribution
where
. Tipping and Bishop prove an analogy of the Gauss-Markov theorem, namely that the affine subspace given by the maximum likelihood estimates of
and
is the PCA subspace (the proof is not difficult but I omit it and refer interested readers to their paper, or Bishop’s Pattern Recognition and Machine Learning book).
It is important to note that although the maximum likelihood estimates of in the pPCA model correspond to the PCA subspace, only posterior distributions can be obtained for the latent data (points on the subspace). Neither the mode nor the mean of those distributions corresponds to the PCA points (orthogonal projections of the observations onto the subspace). However what is true, is that the posterior distributions converge to the PCA points as
. In other words, the relationship between pPCA and PCA is a bit more subtle than that between least squares and regression.
The relationship between regression and (p)PCA is shown in the figure below:
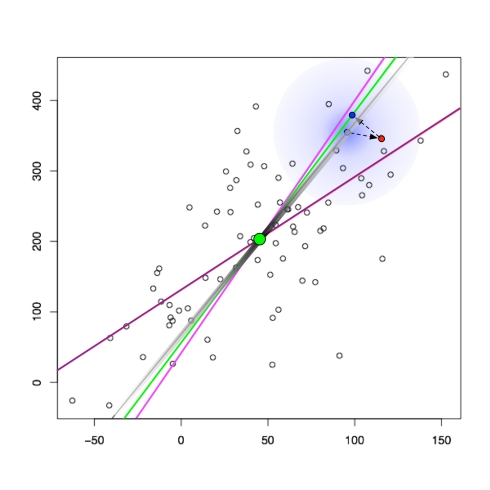 In the figure, points have been generated randomly according to the pPCA model. the black smear shows the affine space on which the points were generated, with the smear indicating the Gaussian distribution used. Subsequently the latent points (light blue on the gray line) were used to make observed points (red) by the addition of isotropic Gaussian noise. The green line is the maximum likelihood estimate for the space, or equivalently by the theorem of Tipping and Bishop the PCA subspace. The projection of the observed points onto the PCA subspace (blue) are the PCA points. The purple line is the least squares line, or equivalently the affine space obtained by regression (y observed as a noisy function of x). The pink line is also a regression line, except where x is observed as a noisy function of y.
In the figure, points have been generated randomly according to the pPCA model. the black smear shows the affine space on which the points were generated, with the smear indicating the Gaussian distribution used. Subsequently the latent points (light blue on the gray line) were used to make observed points (red) by the addition of isotropic Gaussian noise. The green line is the maximum likelihood estimate for the space, or equivalently by the theorem of Tipping and Bishop the PCA subspace. The projection of the observed points onto the PCA subspace (blue) are the PCA points. The purple line is the least squares line, or equivalently the affine space obtained by regression (y observed as a noisy function of x). The pink line is also a regression line, except where x is observed as a noisy function of y.
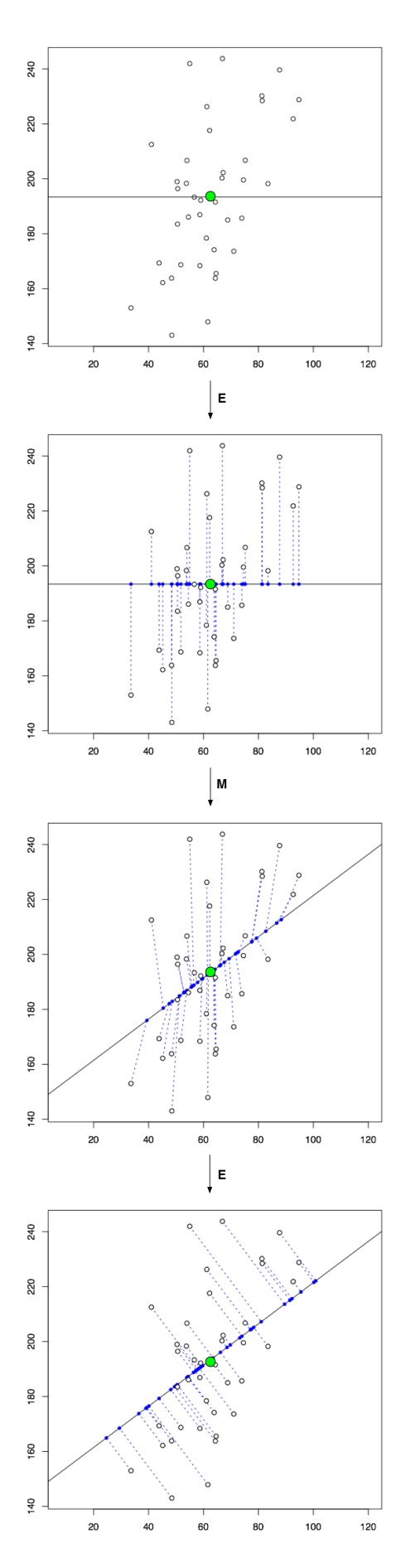
A natural question to ask is why the probabilistic interpretation of PCA (pPCA) is useful or necessary? One reason it is beneficial is that maximum likelihood inference for pPCA involves hidden random variables, and therefore the EM algorithm immediately comes to mind as a solution (the strategy was suggested by both Tipping & Bishop and Roweis). I have not yet discussed how to find the PCA subspace, and the EM algorithm provides an intuitive and direct way to see how it can be done, without the need for writing down any linear algebra:
The exact version of the EM shown above is due to Roweis. In it, one begins with a random affine subspace passing through the centroid of the points. The “E” step (expectation) consists of projecting the points to the subspace. The projected points are considered fixed to the subspace. The “M” step (maximization) then consists of rotating the space so that the total squared distance of the fixed points on the subspace to the observed points is minimized. This is repeated until convergence. Roweis points out that this approach to finding the PCA subspace is equivalent to power iteration for (efficiently) finding eigenvalues of the the sample covariance matrix without computing it directly. This is our first use of the word eigenvalue in describing PCA, and we elaborate on it, and the linear algebra of computing PCA subspaces later in the post.
Another point of note is that pPCA can be viewed as a special case of factor analysis, and this connection provides an immediate starting point for thinking about generalizations of PCA. Specifically, factor analysis corresponds to the model where the covariance matrix
is less constrained, and only required to be diagonal. This is connected to a comment made above about when the PCA subspace might be more useful as a linear fit to data than regression. To reiterate, unlike physics, where some coordinate measurements have very little noise in comparison to others, biological measurements are frequently noisy in all coordinates. In such settings factor analysis is preferable, as the variance in each coordinate is estimated as part of the model. PCA is perhaps a good compromise, as PCA subspaces are easier to find than parameters for factor analysis, yet PCA, via its pPCA interpretation, accounts for noise in all coordinates.
A final comment about pPCA is that it provides a natural framework for thinking about hypothesis testing. The book Statistical Methods: A Geometric Approach by Saville and Wood is essentially about (the geometry of) pPCA and its connection to hypothesis testing. The authors do not use the term pPCA but their starting point is exactly the linear Gaussian model of Tipping and Bishop. The idea is to consider single samples from n independent identically distributed independent Gaussian random variables as one single sample from a high-dimensional multivariate linear Gaussian model with isotropic noise. From that point of view pPCA provides an interpretation for Bessel’s correction. The details are interesting but tangential to our focus on PCA.
We are therefore ready to return to Jeopardy!, where we have “Normality for $600” with the answer “A generalization of linear regression in which the Gaussian noise is isotropic” and the question “What is PCA?”
3. An orthogonal projection of points onto an affine space that maximizes the retained sample variance.
In the previous two interpretations of PCA, the focus was on the PCA affine subspace. However in many uses of PCA the output of interest is the projection of the given points onto the PCA affine space. The projected points have three useful related interpretations:
- As seen in in section 1, the (orthogonally) projected points (red -> blue) are those whose total squared distance to the observed points is minimized.
- What we focus on in this section, is the interpretation that the PCA subspace is the one onto which the (orthogonally) projected points maximize the retained sample variance.
- The topic of the next section, namely that the squared distances between the (orthogonally) projected points are on average (in the
metric) closest to the original distances between the points.
The sample variance of a set of points is the average squared distance from each point to the centroid. Mathematically, if the observed points are translated so that their centroid is at zero (known as zero-centering), and then represented by an matrix X, then the sample covariance matrix is given by
and the sample variance is given by the trace of the matrix. The point is that the jth diagonal entry of
is just
, which is the sample variance of the jth variable. The PCA subspace can be viewed as that subspace with the property that the sample variance of the projections of the observed points onto the subspace is maximized. This is easy to see from the figure above. For each point (blue), Pythagoras’ theorem implies that
. Since the PCA subspace is the one minimizing the total squared red-blue distances, and since the solid black lines (red-green distances) are fixed, it follows that the PCA subspace also maximizes the total squared green-blue distances. In other words, PCA maximizes the retained sample variance.
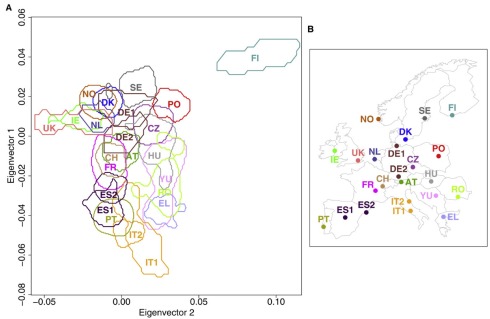
The explanation above is informal, and uses a 1-dimensional PCA subspace in dimension 2 to make the argument. However the argument extends easily to higher dimension, which is typically the setting where PCA is used. In fact, PCA is typically used to “visualize” high dimensional points by projection into dimensions two or three, precisely because of the interpretation provided above, namely that it retains the sample variance. I put visualize in quotes because intuition in two or three dimensions does not always hold in high dimensions. However PCA can be useful for visualization, and one of my favorite examples is the evidence for genes mirroring geography in humans. This was first alluded to by Cavalli-Sforza, but definitively shown by Lao et al., 2008, who analyzed 2541 individuals and showed that PCA of the SNP matrix (approximately) recapitulates geography:
Genes mirror geography from Lao et al. 2008: (Left) PCA of the SNP matrix (2541 individuals x 309,790 SNPs) showing a density map of projected points. (Right) Map of Europe showing locations of the populations .
In the picture above, it is useful to keep in mind that the emergence of geography is occurring in that projection in which the sample variance is maximized. As far as interpretation goes, it is useful to look back at Cavalli-Sforza’s work. He and collaborators who worked on the problem in the 1970s, were unable to obtain a dense SNP matrix due to limited technology of the time. Instead, in Menozzi et al., 1978 they performed PCA of an allele-frequency matrix, i.e. a matrix indexed by populations and allele frequencies instead of individuals and genotypes. Unfortunately they fell into the trap of misinterpreting the biological meaning of the eigenvectors in PCA. Specifically, they inferred migration patterns from contour plots in geographic space obtained by plotting the relative contributions from the eigenvectors, but the effects they observed turned out to be an artifact of PCA. However as we discussed above, PCA can be used quantitatively via the stochastic process for which it solves maximum likelihood inference. It just has to be properly understood.
To conclude this section in Jeopardy! language, we have “Normality for $800” with the answer “A set of points in an affine space obtained via projection of a set of given points so that the sample variance of the projected points is maximized” and the question “What is PCA?”
4. Principal component analysis of Euclidean distance matrices.
In the preceding interpretations of PCA, I have focused on what happens to individual points when projected to a lower dimensional subspace, but it is also interesting to consider what happens to pairs of points. One thing that is clear is that if a pair of points are projected orthogonally onto a low-dimensional affine subspace then the distance between the points in the projection is smaller than the original distance between the points. This is clear because of Pythagoras’ theorem, which implies that the squared distance will shrink unless the points are parallel to the subspace in which case the distance remains the same. An interesting observation is that in fact the PCA subspace is the one with the property where the average (or total) squared distances between the points is maximized. To see this it again suffices to consider only projections onto one dimension (the general case follows by Pythagoras’ theorem). The following lemma, discussed in my previous blog post, makes the connection to the previous discussion:
Lemma: Let be numbers with mean
. If the average squared distance between pairs of points is denoted
and the variance is denoted
then
.
What the lemma says is that the sample variance is equal to the average squared difference between the numbers (i.e. it is a scalar multiple that does not depend on the numbers). I have already discussed that the PCA subspace maximizes the retained variance, and it therefore follows that it also maximizes the average (or total) projected squared distance between the points. Alternately, PCA can be interpreted as minimizing the total (squared) distance that is lost, i,e. if the original distances between the points are given by a distance matrix and the projected distances are given by
, then the PCA subspace minimizes
, where each term in the sum is positive as discussed above.
This interpretation of PCA leads to an interesting application of the method to (Euclidean) distance matrices rather than points. The idea is based on a theorem of Isaac Schoenberg that characterizes Euclidean distance matrices and provides a method for realizing them. The theorem is well-known to structural biologists who work with NMR, because it is one of the foundations used to reconstruct coordinates of structures from distance measurements. It requires a bit of notation: D is a distance matrix with entries and
is the matrix with entries
.
denotes the vector of all ones, and
denotes a vector.
Theorem (Schoenberg, 1938): A matrix D is a Euclidean distance matrix if and only if the matrix is positive semi-definite where
.
For the case when is chosen to be a unit vector, i.e. all entries are zero except one of them equal to 1, the matrix B can be viewed as the Gromov transform (known as the Farris transform in phylogenetics) of the matrix with entries
. Since the matrix B is positive semidefinite it can be written as
, where the matrix X provides coordinates for points that realize D. At this point PCA can be applied resulting in a principal subspace and points on it (the orthogonal projections of X). A point of note is that eigenvectors of
can be computed directly, avoiding the need to compute
which may be a larger matrix if
.
The procedure just described is called classic multidimensional scaling (MDS) and it returns a set of points on a Euclidean subspace with distance matrix that best represent the original distance matrix D in the sense that
is minimized. The term multidimensional scaling without the “classic” has taken on an expanded meaning, namely it encapsulates all methods that seek to approximately realize a distance matrix by points in a low dimensional Euclidean space. Such methods are generally not related to PCA, but classic multidimensional scaling is PCA. This is a general source of confusion and error on the internet. In fact, most articles and course notes I found online describing the connection between MDS and PCA are incorrect. In any case classic multidimensional scaling is a very useful instance of PCA, because it extends the utility of the method to cases where points are not available but distances between them are.
Now we return to Jeopardy! one final time with the final question in the category: “Normality for $1000”. The answer is “Principal component analysis of Euclidean distance matrices” and the question is “What is classic multidimensional scaling?”
An example
To illustrate the interpretations of PCA I have highlighted, I’m including an example in R inspired by an example from another blog post (all commands can be directly pasted into an R console). I’m also providing the example because missing in the discussion above is a description of how to compute PCA subspaces and the projections of points onto them. I therefore explain some of this math in the course of working out the example:
First, I generate a set of points (in ). I’ve chosen a low dimension so that pictures can be drawn that are compatible with some of the examples above. Comments following commands appear after the # character.
set.seed(2) #sets the seed for random number generation. x <- 1:100 #creates a vector x with numbers from 1 to 100 ex <- rnorm(100, 0, 30) #100 normally distributed rand. nos. w/ mean=0, s.d.=30 ey <- rnorm(100, 0, 30) # " " y <- 30 + 2 * x #sets y to be a vector that is a linear function of x x_obs <- x + ex #adds "noise" to x y_obs <- y + ey #adds "noise" to y P <- cbind(x_obs,y_obs) #places points in matrix plot(P,asp=1,col=1) #plot points points(mean(x_obs),mean(y_obs),col=3, pch=19) #show center
At this point a full PCA analysis can be undertaken in R using the command “prcomp”, but in order to illustrate the algorithm I show all the steps below:
M <- cbind(x_obs-mean(x_obs),y_obs-mean(y_obs))#centered matrix MCov <- cov(M) #creates covariance matrix
Note that the covariance matrix is proportional to the matrix $M^tM$. Next I turn to computation of the principal axes:
eigenValues <- eigen(MCov)$values #compute eigenvalues eigenVectors <- eigen(MCov)$vectors #compute eigenvectors
The eigenvectors of the covariance matrix provide the principal axes, and the eigenvalues quantify the fraction of variance explained in each component. This math is explained in many papers and books so we omit it here, except to say that the fact that eigenvalues of the sample covariance matrix are the principal axes follows from recasting the PCA optimization problem as maximization of the Raleigh quotient. A key point is that although I’ve computed the sample covariance matrix explicitly in this example, it is not necessary to do so in practice in order to obtain its eigenvectors. In fact, it is inadvisable to do so. Instead, it is computationally more efficient, and also more stable to directly compute the singular value decomposition of M. The singular value decomposition of M decomposes it into where D is a diagonal matrix and both
and
are orthogonal matrices. I will also not explain in detail the linear algebra of singular value decomposition and its relationship to eigenvectors of the sample covariance matrix (there is plenty of material elsewhere), and only show how to compute it in R:
d <- svd(M)$d #the singular values v <- svd(M)$v #the right singular vectors
The right singular vectors are the eigenvectors of . Next I plot the principal axes:
lines(x_obs,eigenVectors[2,1]/eigenVectors[1,1]*M[x]+mean(y_obs),col=8)
This shows the first principal axis. Note that it passes through the mean as expected. The ratio of the eigenvectors gives the slope of the axis. Next
lines(x_obs,eigenVectors[2,2]/eigenVectors[1,2]*M[x]+mean(y_obs),col=8)
shows the second principal axis, which is orthogonal to the first (recall that the matrix in the singular value decomposition is orthogonal). This can be checked by noting that the second principal axis is also
lines(x_obs,-1/(eigenVectors[2,1]/eigenVectors[1,1])*M[x]+mean(y_obs),col=8)
as the product of orthogonal slopes is -1. Next, I plot the projections of the points onto the first principal component:
trans <- (M%*%v[,1])%*%v[,1] #compute projections of points P_proj <- scale(trans, center=-cbind(mean(x_obs),mean(y_obs)), scale=FALSE) points(P_proj, col=4,pch=19,cex=0.5) #plot projections segments(x_obs,y_obs,P_proj[,1],P_proj[,2],col=4,lty=2) #connect to points
The linear algebra of the projection is simply a rotation followed by a projection (and an extra step to recenter to the coordinates of the original points). Formally, the matrix M of points is rotated by the matrix of eigenvectors W to produce . This is the rotation that has all the optimality properties described above. The matrix T is sometimes called the PCA score matrix. All of the above code produces the following figure, which should be compared to those shown above:
There are many generalizations and modifications to PCA that go far beyond what has been presented here. The first step in generalizing probabilistic PCA is factor analysis, which includes estimation of variance parameters in each coordinate. Since it is rare that “noise” in data will be the same in each coordinate, factor analysis is almost always a better idea than PCA (although the numerical algorithms are more complicated). In other words, I just explained PCA in detail, now I’m saying don’t use it! There are other aspects that have been generalized and extended. For example the Gaussian assumption can be relaxed to other members of the exponential family, an important idea if the data is discrete (as in genetics). Yang et al. 2012 exploit this idea by replacing PCA with logistic PCA for analysis of genotypes. There are also many constrained and regularized versions of PCA, all improving on the basic algorithm to deal with numerous issues and difficulties. Perhaps more importantly, there are issues in using PCA that I have not discussed. A big one is how to choose the PCA dimension to project to in analysis of high-dimensional data. But I am stopping here as I am certain no one is reading at this far into the post anyway…
The take-home message about PCA? Always be thinking when using it!
Acknowledgment: The exposition of PCA in this post began with notes I compiled for my course MCB/Math 239: 14 Lessons in Computational Genomics taught in the Spring of 2013. I thank students in that class for their questions and feedback. None of the material presented in class was new, but the exposition was intended to clarify when PCA ought to be used, and how. I was inspired by the papers of Tipping, Bishop and Roweis on probabilistic PCA in the late 1990s that provided the needed statistical framework for its understanding. Following the class I taught, I benefited greatly from conversations with Nicolas Bray, Brielin Brown, Isaac Joseph and Shannon McCurdy who helped me to further frame PCA in the way presented in this post.





18 comments
Comments feed for this article
May 26, 2014 at 10:04 am
Danny Antaki
Really good break down of PCA
May 26, 2014 at 2:11 pm
pfiziev
Hi Lior,
Great article about PCA!! I spotted two typos that you may wish to correct:
1. In “The generative model provides an interpretation of the independent variables as fixed measured quantities, whereas the dependent variable is a linear combination of the dependent variables with added noise.” The last “dependent” should be “independent”.
and
2. In “Since the PCA subspace is the one minimizing the total squared red-blue distances, and since the solid black lines (red-green distances) are fixed, it follows that the PCA subspace also minimizes the total squared green-blue distances.”, you are actually maximizing the total squared green-blue distances (not minimizing them), because they are equal to d(red,green)^2 – d(red,blue)^2
May 26, 2014 at 3:22 pm
Lior Pachter
Thanks for catching those two typos! They have been fixed.
May 26, 2014 at 3:08 pm
LP
Nice treatment. Hopefully, the population geneticists mentioned early in the post have read the paper that ties PCA together with some more well-known parameters in their field:
“A Genealogical Interpretation of Principal Components Analysis”
Gil McVean
Published: October 16, 2009DOI: 10.1371/journal.pgen.1000686
http://www.plosgenetics.org/article/info%3Adoi%2F10.1371%2Fjournal.pgen.1000686
May 26, 2014 at 3:20 pm
Lior Pachter
Thanks for the link. I really like that paper and the way it connects PCA to the coalescent. My only gripe with it is that PCA is characterized as “explicitly a non-parametric data summary” when in fact the PCA subspaces, by virtue of being equivalent to the pPCA subspaces, make PCA a parametric method involving inference of parameters of a linear Gaussian model (there is also a model selection problem of choosing the dimension).
May 26, 2014 at 3:20 pm
Debora Marks
Thanks Lior. This explains PCA better than any text book I have seen. We should learn to explain all prob and stat methods used in biology this way around, Jeopardy!-rules OK!.
May 27, 2014 at 8:33 am
Robert Flight
pPCA sounds very similar to Peter Wentzell’s mlPCA (maximum likelihood PCA) that uses an alternating least squares algorithm to estimate the subspaces. This is also close to Total Least Squares, and all of these are related to as you said, factor analysis.
I also agree that biologists should be using PCA instead of linear regression due to errors in both X and Y, and many should be using pPCA, mlPCA, and weighted factor analysis methods (weighted alternating least squares, weighted non-negative matrix factorization), but many don’t know that they have non-iid data, or the assumptions made by linear regression or methods like PCA.
May 27, 2014 at 8:35 am
Robert Flight
Forgot to link to the mlPCA paper: http://groupwentzell.chemistry.dal.ca/papers/a34.pdf
And maximum likelyhood weighted alternating least squares: http://groupwentzell.chemistry.dal.ca/papers/a60.pdf
May 31, 2014 at 1:05 pm
R
Just some minor typos, in part 2: the Gaussian distribution should be noted f(x, \mu \sigma), and, in the following expressions, e.g. L(\mu, \sigma), x should probably be x_i.
May 31, 2014 at 1:08 pm
Lior Pachter
Thanks!
May 31, 2014 at 9:23 pm
dpoznik
Great post. Spotted what I believe to be a couple minor typos:
1. In “Note that the covariance matrix is the matrix M’M,” I would replace “M’M” with “M’M/(n-1)”. Or perhaps just add “proportional to.”
2. I think this equation “\tilde{X} = min_{M \mbox{ of rank } k} ||X-M||_2” should be an “arg min” rather than a “min.” Also, the equation has rendered with a spurious “\hat{A}” after the “f”.
May 31, 2014 at 9:47 pm
Lior Pachter
Thanks for reading carefully! I’ve corrected the typos including some others spotted by critical readers. Thanks to all of you.
June 2, 2014 at 1:27 am
Erik van Nimwegen
Hi Lior,
very nice post. I did not see you mention what I consider to be the simplest and most transparent probabilistic interpretation of PCA. That is, assume your n datapoint from R^p derive from a multi-variate Gaussian in p dimensions. The maximum likelihood estimate of this multi-variate Gaussian has mean equal to the sample mean and a covariance matrix equal to the sample covariance matrix (note: with factors 1/n, not 1/(n-1)).
PCA now simply consists in
1. Diagonalizing this covariance matrix. That is, to find the linear combinations of the original p variables that are statistically independent (in the ML estimate).
2. Decide to approximate the original multi-variate Gaussian by a lower-dimensional one that only retains the first k<p eigenvectors.
In other words, one get the ML multi-variate Gaussian from the data, rotates to a basis in which fluctuations along each axis are independent, and then retains only the axes with the highest variance. That's how I present it to my students.
btw. Although I must admit I have not studied the pPCA papers, it would seem to me that that interpretation is subsumed under the simple interpretation above (the final covariance matrix is simply the sum of the covariance matrix W on the subspace plus the diagonal matrix sigma^2 I).
June 2, 2014 at 11:01 pm
oz
Thanks a lot for a great post clarifying many points about PCA.
Some minor comments which can improve even further (at least my) understanding:
1. In the pPCA interpretation, you write down the generative model
for the observed t. If the \epsilon is additive Gaussian noise, then why is \psi needed? why not simply write
t ~ N(\mu, W W^T + \sigma^2 I) ?
Also, you didn’t write a generative model for the hidden variables x.
Is x simply a multi-variate Gaussian with mean zero and unit covariance matrix? x ~ N(0, I) ?
2. In Schoenberg’s theorem, there is a quantifier missing so I’m not sure what the Thm. says. I assume that the statement is:
‘.. if there exists s with s’1=1, such that .. is positive semidefinite’ and not ‘for all s such that s’1=1, we have … is positive semi-definite’. Is that correct?
3. You write that classic MDS is obtained when taking s to be a unit vector . But doesn’t this involve choosing which unit vector to use? (where the ‘1’ in one of the coordinates corresponds to one of the data points). I think of MDS as symmetric with respect to the data points, and this choice of s treats one of them differently – or am I missing something here and the choice of s doesn’t matter?
4. You did not write about the computational complexity of computing PCA.
As far as I understand, computing the full singular-value decomposition requires O(n^3) computations, but if you are interested only in the few top eigenvectors and eigenvalues, there are faster (linear?) algorithms. Is it possible to elaborate/give pointers to that?
December 5, 2015 at 1:11 am
daniel
Just stumbled on this and really clarified. Showing my CS189 (Machine Learning) friends 😉
July 25, 2016 at 10:52 am
NameReq
Nice explanation, thanks. I am a bit confused to your reference to an orthogonal (orthonormal?) set of vectors “induced” by the flag. AFAIK, the flag is a nested collection of affine spaces; I don’t see how a flag gives rise to a collection of vectors. Would you explain?
Thanks.
July 25, 2016 at 10:54 am
NameReq
Hi, me again. Sorry, just to note that the point I am referring to is in the paragraph immediately below your point 3 above.
September 8, 2018 at 12:04 pm
steven gortler
In the post, you said “Alternately, PCA can be interpreted as minimizing the total (squared) distance that is lost”. That is clearly true if we assume that we will be obtaining our $\tilde{x}$ through an orthogonal projection onto some affine space (Pythagoras).
Is it still true if we drop that assumption, and minimize over all low dimensional embeddings of the points? (I think it is not,
and this corresponds to the idea that “classical scaling MDS”/PCA, optimizes the so called “STRAIN” energy, but not the “SSTRESS” energy.)