I recently published a paper on the bioRxiv together with Vasilis Ntranos, Lynn Yi and Páll Melsted on Identification of transcriptional signatures for cell types from single-cell RNA-Seq. The contributions of the paper can be summed up as:
- The simple technique of logistic regression, by taking advantage of the large number of cells assayed in single-cell RNA-Seq experiments, is much more effective than current approaches at identifying marker genes for clusters of cells.
- The simplest single-cell RNA-Seq data, namely 3′ single-end reads produced by technologies such as Drop-Seq or 10X, can distinguish isoforms of genes.
- The simple idea of GDE provides a unified perspective on DGE, DTU and DTE.
These simple, simple and simple ideas are so obvious that of course anyone could have discovered them, and one might be tempted to go so far as to say that even if people didn’t explicitly write them down, they were basically already known. After all, logistic regression was published by David Cox in 1958, and who didn’t know that there are many 3′ unannotated UTRs in the human genome? As for DGE, DTU and DTE (and DTE->G and DTE+G) I mean who doesn’t get these basic concepts? Indeed, after reading our paper someone remarked that one of the key results “was already known“, presumably because the successful application of logistic regression as a gene differential expression method for single-cell RNA-Seq follows from the fact that Šidák aggregation fails for differential gene expression in bulk RNA-Seq.
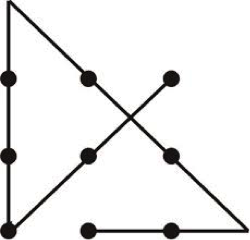
The “was already known” comment reminded me of a recent blog post about the dirty secret of mathematics. In the post, the author begins with the following math problem: Without taking your pencil off the paper/screen, can you draw four straight lines that go through the middle of all of the dots?

The problem may not yield immediately (try it!) but the solution is obvious once presented. This is a case of the solution requiring a bit of out-of-the-box thinking, leading to a perspective on the problem that is obvious in retrospect. In the Ntranos, Yi et al. paper, the change in perspective was the realization that “Instead of the traditional approach of using the cell labels as covariates for gene expression, logistic regression incorporates transcript quantifications as covariates for cell labels”. It’s no surprise the “was already known” reaction reared it’s head in this case. It’s easy to convince oneself, after the fact, that the “obvious” idea was in one’s head all along.
The egg of Columbus is an apocryphal tale about ideas that seem trivial after the fact. The story originates from the book “History of the New World” by Girolamo Benzoni, who wrote that Columbus, upon upon being told that his journey to the West Indies was unremarkable and that Spain “would not have been devoid of a man who would have attempted the same” had he not undertaken the journey, replied
“Gentlemen, I will lay a wager with any of you, that you will not make this egg stand up as I will, naked and without anything at all.” They all tried, and no one succeeded in making it stand up. When the egg came round to the hands of Columbus, by beating it down on the table he fixed it, having thus crushed a little of one end”

The story makes a good point. Discovery of the Caribbean in the 6th millennium BC was certainly not a trivial accomplishment even if it was obvious after the fact. The egg trick, which Columbus would have learned from the Amerindians who first brought chickens to the Americas, is a good metaphor for the discovery.
There are many Amerindian eggs in mathematics, which has its own apocryphal story to make the point: A professor proving a theorem during a lecture pauses to remark that “it is obvious that…”, upon which she is interrupted by a student asking if that’s truly the case. The professor runs out of the classroom to a nearby office, returning after several minutes with a notepad filled with equations to exclaim “Why yes, it is obvious!” But even first-rate mathematicians can struggle to accept Amerindian eggs as worthy contributions, frequently succumbing to the temptation of dismissing others’ work as obvious. One of my former graduate school mentors was G.W. Peck, a math professor who created a pseudonym for the express purpose of publishing his Ameridian eggs in a way that would reduce unintended embarrassment for those whose work he was improving on in in “trivial ways”. G.W. Peck has an impressive publication record.
Bioinformatics is not very different from mathematics; the literature is populated with many Amerindian eggs. My favorite example is the Smith-Waterman algorithm, an algorithm for local alignment published by Temple Smith and Michael Waterman in 1981. The Smith-Waterman algorithm is a simple modification of the Needleman-Wunsch algorithm:

The table above shows the differences. That’s it! This table made for a (highly cited) paper. Just initialize the Needleman-Wunsch algorithm with zeroes instead of a gap penalty, set negative scores to 0, trace back from the highest score. In fact, it’s such a minor modification that when I first learned the details of the algorithm I thought “This is obvious! After all, it’s just the Needleman-Wunsch algorithm. Why does it even have a name?! Smith and Waterman got a highly cited paper?! For this?!” My skepticism lasted only as long as it took me to discover and read Peter Sellers’ 1980 paper attempting to solve the same problem. It’s a lot more complicated, relying on the idea of “inductive steps”, and requires untangling mysterious diagrams such as:

The Smith-Waterman solution was clever, simple and obvious (after the fact). Such ideas are a hallmark of Michael Waterman’s distinguished career. Consider the Lander-Waterman model, which is a formula for the expected number of contigs in a shotgun sequencing experiment:
Here N is the number of reads sequenced and R=NL/G is the “redundancy” (reads * fragment length / genome length). At first glance the Lander-Waterman “model” is just a formula arising from the Poisson distribution! It was obvious… immediately after they published it. The Pevzner-Tang-Waterman approach to DNA assembly is another good example. It is no coincidence that all of these foundational, important and impactful ideas have Waterman in their name.
Looking back at my own career, some of the most satisfying projects have been Amerindian eggs, projects where I was lucky to participate in collaborations leading to ideas that were obvious (after the fact). Nowadays I know I’ve hit the mark when I receive the most authentic of compliments: “your work is trivial!” or “was widely known in the field“, as I did recently after blogging about plagiarism of key ideas from kallisto. However I’m still waiting to hear the ultimate compliment: “everything you do is obvious and was already known!”
(Click “read the rest of this entry” to see the solution to the 9 dot problem.)

6 comments
Comments feed for this article
February 15, 2018 at 5:53 pm
Sandra
I appreciate what you have written, because there is elegance in showing these results. However, I think these results also beg some deeper questions within the scientific community. For example, maybe some scientists would never bother to publish such results because they do not feel they are enough of a contribution? Or perhaps we need to ask, would such results even be published if the authors’ names were Jane Doe or Juan Vegas?
I think implicit bias may be a factor too. Would Jane Doe or Juan Vegas have such a work rejected because it is too trivial?
February 16, 2018 at 8:18 am
Leonardo de Oliveira Martins
My favourite example is the Lineweaver–Burk equation, which is a “trivial inversion” of the Michaelis–Menten equation.
March 1, 2018 at 2:21 am
fedster9
“The egg trick, which Columbus would have learned from the Amerindians who first brought chickens to the Americas, is a good metaphor for the discovery.” How do we know Columbus learned said trick from someone? if he did learn it from someone how do we know it was Amerindian? how do we know Amerindians used this trick? These are far from trivial questions!
December 6, 2018 at 4:44 am
Aubrey Bailey
“RSEM17 was used to generated uniformly”
January 2, 2019 at 7:54 pm
vasilispromponas
Temple Smith described during an ISMB keynote talk (in Vienna, 2007 if I correctly recall) their efforts to publish the S-W paper in one of the high-impact journals and getting rejected. They published their gem in JMB. Simple, elegant, inspiring work!!
October 24, 2019 at 1:34 pm
Zach
I wanted to drop a thank you note for this blog post! I had a lot of trouble pushing my first publication out, as I felt it was obvious and trivial (although not previously reported). After the first publication, we started a subgroup to further develop this method, and grew from 1 to 5 in the first couple months, not to mention receiving many grants to support the follow-up work (and personnel). This post was very encouraging, and I still come back to read it from time to time.