[Update April 6, 2014: The initial title of this post was “23andme genotypes are all wrong”. While that was and remains a technically correct statement, I have changed it because the readership of my blog, and this post in particular, has changed. Initially, when I made this post, the readers of the blog were (computational) biologists with extensive knowledge of genotyping and association mapping, and they could understand the point I was trying to make with the title. However in the past few months the readership of my blog has grown greatly, and the post is now reaching a wide public audience. The revised title clarifies that the content of this post relates to the point that low error rates in genotyping can be problematic in the context of genome-wide association reports because of multiple-testing.]
I have been reading the flurry of news articles and blog posts written this week about 23andme and the FDA with some interest. In my research talks, I am fond of displaying 23andme results, and have found that people always respond with interest. On the teaching side, I have subsidized 23andme testing for volunteer students in Math127 who were interested in genetics so that they could learn about personalized genomics first-hand. Finally, a number of my former and current students have worked at 23andme, and some are current employees.
Despite lots of opinions being expressed about the 23andme vs. FDA kerfuffle, I believe that two key points have been ignored in the discussions:
- All 23andme genotypes that have ever been reported to customers are wrong. This is the case despite very accurate genotyping technology used by 23andme.
- The interpretation of 23andme results involves examining a large number of odds ratios. The presence of errors leads to a huge multiple-testing problem.
Together, these issues lead to an interesting conundrum for the company, for customers, and for the FDA.
I always find it useful to think about problems concretely. In the case of 23andme, it means examining actual genotypes. Fortunately, you don’t have to pay the company $99 dollars to get your own- numerous helpful volunteers have posted their 23andme genotypes online. They can be viewed at openSNP.org where “customers of direct-to-customer genetic tests [can] publish their test results, find others with similar genetic variations, learn more about their results, get the latest primary literature on their variations and help scientists find new associations”. There are a total of 624 genotypes available at openSNP, many of them from 23andme. As an example, consider “samantha“, who in addition to providing her 23andme genotype, also provides lots of phenotypic information. Here is the initial part of her genotype file:
# This data file generated by 23andMe at: Wed Jul 20 20:37:11 2011 # # Below is a text version of your data. Fields are TAB-separated # Each line corresponds to a single SNP. For each SNP, we provide its identifier # (an rsid or an internal id), its location on the reference human genome, and the # genotype call oriented with respect to the plus strand on the human reference # sequence. We are using reference human assembly build 36. Note that it is possible # that data downloaded at different times may be different due to ongoing improvements # in our ability to call genotypes. More information about these changes can be found at: # https://www.23andme.com/you/download/revisions/ # # More information on reference human assembly build 36: # http://www.ncbi.nlm.nih.gov/projects/mapview/map_search.cgi?taxid=9606&build=36 # # rsid chromosome position genotype rs4477212 1 72017 AA rs3094315 1 742429 AG rs3131972 1 742584 AG rs12124819 1 766409 AA rs11240777 1 788822 AA rs6681049 1 789870 CC rs4970383 1 828418 CC rs4475691 1 836671 CC rs7537756 1 844113 AA rs13302982 1 851671 GG rs1110052 1 863421 GT ...
Anyone who has been genotyped by 23andme can get this file for themselves from the website (by clicking on their name, then on “Browse Raw Data” from the pull-down menu, and then clicking on “Download” in the top-right corner of the browser window). The SNPs are labeled with rsid labels (e.g. rs3094315) and correspond to specific locations on chromosomes (e.g. chr1:742429). Since every human is diploid, two bases are shown for every SNP; one came from mom and one from dad. The 23andme genotype is not phased, which means that you can’t tell in the case of rs3094315 whether the A was from mom and the G from dad, or vice versa (it turns out paternal origin can be important, but that is a topic for another post).
A key question the FDA has asked, as it does for any diagnostic test, is whether the SNP calls are accurate. The answer is already out there. First, someone has performed a 23andme replicate experiment precisely to assess the error rate. In an experiment in 2010 with two replicates, 85 SNPs out of about 600,000 were different. Today, Illumina types around 1 million SNPs, so one would expect even more errors. Furthermore, a replicate analysis provides only a lower bound, since systematic errors will not be detected. Another way to examine the error rate is to look at genotypes of siblings. That was written about in this blog post which concluded there were 87 errors. 23andme currently uses the Illumina Omni Express for genotyping, and the Illumina spec sheet claims a similar error rate to those inferred in the blog posts mentioned above. The bottom line is that even though the error rate for any individual SNP call is very very low (<0.01% error), with a million SNPs being called there is (almost) certainly at least one error somewhere in the genotype. In fact, assuming a conservative error rate leading to an average of 100 errors per genotype, the probability that a 23andme genotype has no errors is less than 10^(-40).
The fact that 23andme genotypes are wrong (i.e. at least one error in some SNP) wouldn’t matter if one was only interested in a single SNP. With very high probability, it would be some other SNPs that are the wrong ones. But the way people use 23andme is not to look at a single SNP of interest, but rather to scan the results from all SNPs to find out whether there is some genetic variant with large (negative) effect. The good news is that there isn’t much information available for the majority of the 1 million SNPs being tested. But there are, nevertheless, lots of SNPs (thousands) to look at. Whereas a comprehensive exam at a doctor’s office might currently constitute a handful of tests– a dozen or a few dozen at most– a 23andme test assessing thousands of SNPs and hundreds of diseases/traits constitutes more diagnostic tests on an individual at one time than have previously been performed in a lifetime.
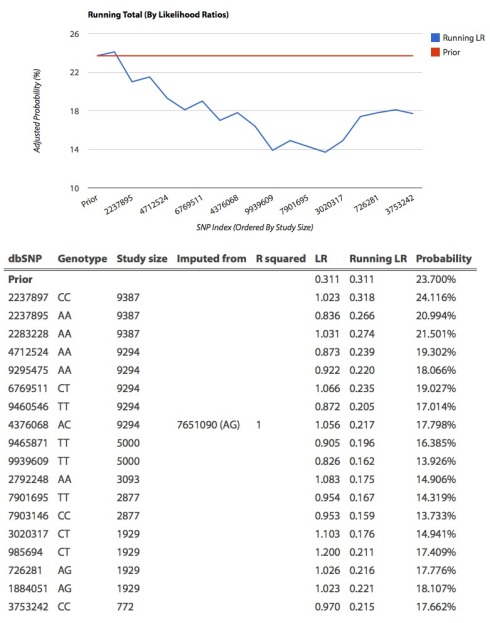
To understand how many tests are being performed in a 23andme experiment, it is helpful to look at the Interpretome website. The website allows a user to examine information on SNPs without paying, and without uploading the data. I took a look at Samantha, and the Interpretome gave information about 2829 SNPs. These are SNPs for which there is a research article that has identified the SNP as significant in some association study (the website conveniently provides direct links to the articles). For example, here are two rows from the phenotype table describing something about Samantha’s genetic predisposition for large head circumference:
Head circumference (infant) 11655470 CC T .05 4E-6 22504419
Head circumference (infant) 1042725 CC T .07 3E-10 22504419
Samantha’s genotype at the locus is CC, the “risk” allele is T, the odds ratios are very small (0.05,0.07) and the p-values are apparently significant. Interpretome’s results differ from those of 23andme, but looking at the diversity of phenotypes reported on gives one a sense for the possibilities that currently exist in genetics, and the scope of 23andme’s reports.
From the estimates of error rates provided above, and using the back of an envelope, it stands to reason that about 1/3 of 23andme tested individuals have an error at one of their “interesting” SNPs. Not all of SNPs arising in association studies are related to diseases, but many of them are. I don’t think its unreasonable to postulate that a significant percentage of 23andme customers have some error in a SNP that is medically important. Whether such errors are typically false positives or false negatives is unclear, and the extent to which they may lead to significant odds ratios is another interesting question. In other words, its not good enough to know how frequently warfarin sensitivity is being called incorrectly. The question is how frequently some medically significant result is incorrect.
Of course, the issue of multiple testing as it pertains to interpreting genotypes is probably a secondary issue with 23andme. As many bloggers have pointed out, it is not even clear that many of 23andme’s odds ratios are accurate or meaningful. A major issue, for example, is the population background of an individual examining his/her genotype and how close it is to the population on which the GWAS were performed. Furthermore, there are serious questions about the meaning of the GWAS odds ratios in the case of complex traits. However I think the issue of multiple testing is a deeper one, and a problem that will only be exacerbated as more disease SNPs are identified. Having said that, there are also approaches that could mitigate errors and improve fidelity of the tests. As DECODE genetics has demonstrated, imputation and phasing can in principle be used to infer population haplotypes, which not only are useful for GWAS analyses, but can also be used to identify erroneous SNP calls. 23andme’s problem is that although they have many genotypes, they are from diverse populations that will be harder to impute and phase.
The issue of multiple testing arising in the context of 23andme and the contrast with classic diagnostics reminds me of the dichotomy between whole-genome analysis and classic single gene molecular biology. The way in which customers are looking at their 23andme results is precisely to look for the largest effects, i.e. phenotypes where they appear to have high odds of contracting a disease, or being sensitive to some drug. This is the equivalent of genome scientists picking the “low hanging fruit” out of genome-wide experiments such as those performed in ENCODE. In genomics, scientists have learned (with some exceptions) how to interpret genome-wide analyses after correcting for multiple-hypothesis testing by controlling for false discovery rate. But are the customers of 23andme doing so? Is the company helping them do it? Should it? Will the FDA require it? Can looking at ones own genotype constitute too much testing?
There are certainly many precedents for superfluous harmful testing in medicine. For example, the American Academy of Family Physicians has concluded that prostate cancer PSA tests and digital rectal exams have marginal benefits that are outweighed by the harm caused by following up on positive results. Similar arguments have been made for mammography screening. I therefore think that there are serious issues to consider about the implications of direct-to-consumer genetic testing and although I support the democratization of genomics, I’m glad the FDA is paying attention.
Samantha’s type 2 diabetes risk as estimated from her genotype by Interpretome. She appears to have a lower risk than an average person. Does this make it ok for her to have another cookie?

35 comments
Comments feed for this article
November 30, 2013 at 11:53 am
cooplab
There are obviously multiple ways to look at this problem.
In my view the error rate vs the true positive rate at SNPs with serious consequences/predictive ability is the important thing. With an error rate as low as 23&me’s the prediction of an serious allele at a SNP is more likely a true positive than a false positive in the frequency of the allele in the population is >1/10000. While if there are large numbers of SNPs with serious consequences I may have a few false positives I will have also learnt medically important, and true, information at many more actionable loci. That seems like the important thing.
If memory serves 23&me suggest that you followup with a genetic counselor on serious predictions, who will get you retested. The FDA should be making sure that this information is reported in the responsible way, and the community should be helping them do this. However, reporting genome-wide error rates, not accounting for the rate of true positives doesn’t seem like a particularly useful way to discuss the issues at hand.
November 30, 2013 at 12:13 pm
Lior Pachter
I completely agree with you. If you read my post I never said the point was for 23andme to report genomewide error rates. I was making the case that it’s important to understand the balance between true and false positives, that it is not entirely trivial to do so, and that 23andme should be required to evaluate and disclose accuracy.
Of course one can and always should retest. And of course genetic information is useful. But 23andme is marketing to millions potential customers and the fact that the FDA is serious about regulating them is good.
November 30, 2013 at 12:38 pm
cooplab
Lior,
I agree that there are subtle points here, and as I said there are various perspectives. I agree it is good that the FDA is looking into it. I did read your post. Throughout the post you repeatedly talk about the chance that a person has one inaccurate result reported, and you mention that the problem getting worse the more associations are found. That does seem to be written in a way that does not fully acknowledge the fp/tp rates, and certainly in a way that will likely confuse many people.
Graham
November 30, 2013 at 12:08 pm
Dan Reghecampf
It is stated that: “Since every human is diploid, two bases are shown for every SNP; one can from mom and one from dad.”
That is not entirely true! Always there are exceptions in biology! For example, males have in majority of cases have only one chromosome Y and only one chromsome X.
For example, this can be easily seen looking to the genotype data of Alexandre Bolze (see here: http://opensnp.org/users/1401) which shows like this:
…
rs10156975 X 85124905 G
rs3790357 X 85125119 A
i5003829 X 85134049 G
rs719988 X 85148366 G
rs16980331 X 85156911 T
…
rs13447361 Y 2821786 G
rs2267801 Y 2828196 T
rs2267802 Y 2828425 A
rs9786142 Y 2842212 A
…
i3001860 MT 16338 A
i4000563 MT 16339 C
i4000564 MT 16340 A
i3001861 MT 16342 T
…
November 30, 2013 at 12:11 pm
Michael Eisen
Multiple testing is a problem. But it’s a problem of there being lots of possible tests, not that one company happens to be doing them all. If the tests for individual SNPs, phenotypes, have an acceptably low error rate if done in isolation, then it doesn’t make sense to say that 23andme is somehow problematic because they are doing lots of tests.
November 30, 2013 at 12:24 pm
Lior Pachter
Mike, all I said was that I’m glad the FDA is paying attention. Not specifically that they are paying attention to 23andme.
My post is about 23andme’s error rates because they are the only company (I think) offering consumers direct access and interpretation of Illumina’s gentotyping.
December 18, 2015 at 12:40 pm
Chris
I got tested… My grandmother came straight from Italy with direct knowledge of hundreds of years of Italians so how am I only 3% or 3% of my Dna?
November 30, 2013 at 12:17 pm
Dan Reghecampf
Regarding this “In other words, its not good enough to know how frequently warfarin sensitivity is being called incorrectly.”
It just has been published a scientific study, that is E.S. Kimmel et al. “A Pharmacogenetic versus a Clinical Algorithm for Warfarin Dosing”, The New England Journal of Medicine, November 19, 2013, DOI: 10.1056/NEJMoa1310669, , which shows that actually that there is no connection between genotype and the sensitivity to Wafarin (that is anticoagulation). The conclusion from the above study is a based on a much larger clinical trials (~ 1015 patients) than previously!
The previous studies, where this link was found, were based on very small clinical trials or observational studies (i.e. smaller than the previously mentioned one).
December 6, 2013 at 12:32 am
Roxana Daneshjou (@RoxanaDaneshjou)
If you read all three papers, you will notice a few things:
-Kimmel et al. compares a clinical algorithm (not standard of care currently) vs. a clinical + PgX algorithm that only uses genotyping data for the revision algorithm, claiming that it’s not useful in the initial dosing. No difference is seen in time spent in therapeutic INR over 4 weeks.
-Verhoef uses a different clinical (still not standard of care) vs. PgX + clinical algorithm. Finds a difference in time spent in therapeutic INR (PgX + clinical algorithm does better) during the first 4 weeks. No difference over 12 weeks.
-Pirmohamed uses current standard of care vs. the IWPC PgX dosing algorithm. PgX algorithm wins in time spent to INR (this doesn’t answer if it’s the clinical or the PgX part that is better, but in the original IWPC paper, the PgX algorithm was definitely better than the clinical only retrospectively).
My point: You can’t claim based on these findings that genotype has no connection to warfarin sensitivity. Those studies really focused on time in therapeutic INR, and the studies were underpowered to look at some secondary outcomes (Kimmel et al. says that it was underpowered to look at one of the most important outcomes –bleeding events). Moreover, each study used a different algorithm, did not take into account other variants besides the main three, and definitely did not include rare variation.
December 6, 2013 at 6:26 pm
Darya Filippova
As I recall, in Pirmohamed paper they claim a difference of about 6% between standard of care vs. IWPC algorithm — which may still fail to justify the expense of genotyping. It would be interesting to see a follow up study that looks at #bleeding or thrombosis events though.
November 30, 2013 at 12:31 pm
Yaniv Erlich
Lior,
I am affraid that this tine you got it upside down. The absolute noise level (probability of an error) is not interesting and misleading. From a statistical point of view, we should focus on the area under the curve. This number should get better as *more* SNP are associated with a disease. Think how many errors a blood panel has. According to your logic, we should go with a very narrow panel to reduce the noise. In fact, we should do zero test because than the probability of an error is zero! Well, I prefer to be vaguely right than precisely wrong….
November 30, 2013 at 12:33 pm
Lior Pachter
Yaniv,
The issue is not how many SNPs associate with a disease, it’s how many diseases are being tested. If we were testing one disease with the whole genome you would be correct. But the number of phenotypes being tested is growing and my point is simply that this is an issue that needs to be considered.
November 30, 2013 at 3:57 pm
Ruchira Datta
Another issue may be that the risky variant is linearly associated with several SNPs.
For instance, suppose SNP A at position x is the causative SNP for an increased risk of some disease, say making it 10% more likely. During crossover, DNA from pairs of chromosomes is recombined. The closer together two positions are on a chromosome, the less likely they will be split by recombination, i.e., the more likely that they will be inherited together. So, it could be that SNP B at position y which is very close to position x, has nothing intrinsically to do with the disease. However, SNP B is likely to be inherited together with SNP A. So, if we just look at SNP B alone, it seems that it increases the risk of the same disease by, say, 8%. This is actually a product of two factors: i) the increased likelihood that one has SNP A given that one has SNP B, and ii) the increased likelihood that one has the disease, given that one has SNP A.
Now, suppose we do a test and find that the person has both SNP A and SNP B. If we just naively multiply the increased risks together, we get that the person has an 18.8% higher risk of the disease. But this would be wrong. We actually measured SNP A. Once we measured that, SNP B gives us no new information about the likelihood that one has SNP A. The increased risk of the disease is just 10%, coming from the causative SNP, i.e., SNP A.
The situation is more complicated because we don’t actually know “the causative SNP”, and many diseases may have multiple SNPs contributing causally. Of course there are techniques to deal with all these uncertainties. However those techniques have to be evaluated carefully.
November 30, 2013 at 4:04 pm
Ruchira Datta
Actually, if we take into account the possibility of genotyping errors, measuring both SNP A and SNP B together does yield increased confidence that we measured SNP A. However, this is a much smaller effect; the numbers I wrote above assume no genotyping errors.
November 30, 2013 at 4:41 pm
Nick Eriksson
Lior,
Of course, the error rate for genotyping is not anywhere near uniform. In particular, the SNPs in reports on the 23andMe website will tend to be much better than random SNPs (for a variety of reasons). Such well performing probes will have miscall rates of essentially zero (with nocall rates around 1/1000 or less), so “about 1/3 of 23andme tested individuals have an error at one of their ‘interesting’ SNPs” is not very accurate.
And of course, different errors matter in different ways. It’s pretty amusing that of the two SNPs from intrepretome you cite, there is a 50% false positive rate – a SNP with a p-value of 4E-6 is almost certainly not a true signal. But for a small effect for a trait with lots of SNPs – an error just doesn’t matter.
For multiple testing, you don’t quite spell out the exact problem. Perhaps what you’re saying is that if a customer receives 40 risk reports, she will almost certainly be in the top 5% of people for one of these reports. This is an interesting point – perhaps instead of the typical question of “what is the AUC achieved for one disease”, one should be asking “what is the probability that you will get one of your top two predicted diseases”.
Which is nice, because predictive accuracy for “chance you will get one or more of the diseases you are in the 95% risk percentile for” will be better than it would be for a single disease, as long as this interpretation is limited to common diseases.
Granted, if 23andMe started providing risk for more and more (rarer and rarer) diseases, this could be a problem. If I were to rank the top 40 astroids that might fall on your house, it doesn’t make much sense to worry about the top two. But I’m not sure there will be that many large GWAS of very rare diseases, so…
-Nick
November 30, 2013 at 6:05 pm
Lior Pachter
Nick,
Thanks for your thoughtful response. I’d like to follow up briefly on a few points you make.
First, I am happy to hear that 23andme is more accurate on reported SNPs than on an average SNP call. Hopefully that will go a long way to allay the issues with the FDA. As you know, I’m very supportive of the mission of democratizing genetics (as I stated in my post) and I’d like nothing more than to see 23andme lead the way in addressing issues about genetics for the public.
The points I was raising about accuracy and multiple testing however have to do with the issue that a standard Illumina error rate can be unacceptable when looking at many SNPs. Although my number of 3000 that I used for “interesting” is high– there are far fewer actionable SNPs today– I don’t think it is a stretch in the (near) future. My point was merely that issues of accuracy can be subtle when looking at multiple phenotypes, and that I think its good to have regulatory oversight because of this. For example, accuracy could easily be improved at every SNP by spending more money, simply by performing replicates. However the extent to which this is necessary is exactly why we have regulatory oversight for medical diagnostics.
This issue is not specific to 23andme. In fact, your point about the Interpretome SNP I highlighted makes the case that Interpretome should probably be regulated as well!
You raise a separate issue about the multiple testing issue arising from looking at multiple test reports. Thats a very interesting point, which I should have addressed but didn’t- thanks for bringing it up. I agree that predictive accuracy will be higher for “chance you will get one or more diseases you are in the 95% risk percentile for”, but then again, such a result requires what I think is fairly characterized as subtle statistical understanding by a lay person. Again, I think its good that the FDA should regulate the testing and reporting for such results.
Thanks again,
Lior
November 30, 2013 at 8:29 pm
Obi Griffith
Nice post Lior. Agree with everything you’ve said. I personally feel that the FDA (and certainly much of the coverage) is over-reacting. But agree these are important issues. I was interested to read about that two replicate experiment. “In an experiment in 2010 with two replicates, 85 SNPs out of about 600,000 were different. Today, Illumina types around 1 million SNPs, so one would expect even more errors.” To add another data point. My twin and I both did 23andMe in Apr 2011. It was with the 1million SNP chip (actually ~930,000) I think). There were 3242 differences. But most of these were cases where one or both results failed to make a call. If (as in the blog you link to) we exclude “no calls” then we observed only 64 differences. So actually, that is better than the previous (85 differences) result despite approximately 50% more total SNPs and genotyping two different people (albeit with “identical” genomes). Perhaps they new kits have lower error rate or they’ve improved the software. This is actually how we discovered we are identical twins. It turns out that, without genetic testing (which was/is not standard), misdiagnosing identical vs fraternal twins is actually surprisingly common. So, that alone was worth the $100!
November 30, 2013 at 8:45 pm
Kevin McLoughlin
I had myself and some friends genotyped by 23andMe a few months ago, and didn’t find anything too interesting in the various “health reports” they provide. So I downloaded the raw SNP data and wrote some code to match my and my friends’ alleles against the risk alleles identified in all GWAS studies published to date (NIH maintains a database summarizing these). Not surprisingly, there were lots of “hits” indicating higher risk for dozens of different disorders. However, after looking at the data for a while, I realized that these results – and nearly *every* other prediction of disease risk based on GWAS studies – simply don’t mean that much.
It’s not just the multiple testing problem, though that’s always an issue whenever you’re fishing for something “interesting” rather than testing for a specific disease. The big issues with predictions based on GWAS studies are that (1) the relative risk factors for *most* diseases for *most* individual SNPs are tiny, on the order of 1.1X to 1.7X; (2) the risk alleles are almost always common; it’s unusual for them to have allele frequencies below 10%; (3) as Lior pointed out, the relative risk is highly dependent on the background population; and (4) there’s typically no attempt to look for epistatic interations between different SNP loci. For some of the 23andMe health reports, they do look at multiple SNP loci, but I believe they’re just multiplying the relative risks or the odds ratios, as if there were no epistasis, which I strongly suspect is wrong for most of the diseases they’re reporting on.
This doesn’t mean I necessarily think it’s good that the FDA is trying to regulate DTC genetic testing, though. Let’s just say I have mixed feelings about it, given what I know about how the FDA works…
November 30, 2013 at 9:29 pm
Simon
Many CLIA labs are using such arrays for clinical carrier screening, many confirm positive genotypes by sanger – not all, in clinical labs the accuracy is never 100% – no matter what the lab will tell you.
FDA approved kits are never 100%, this is a fact of life.
December 1, 2013 at 5:22 am
Mark James Adams (@mja)
I’m trying to get a handle on Lior’s assertion that “it stands to reason that about 1/3 of 23andme tested individuals have an error at one of their “interesting” SNPs.”
Taking the harmonic mean of the mistyping rates that I’ve seen in this post and the comments yields 7e-5. I am assuming this is the mistyping rate of the genotype, not each allele at each loci.
So if we have 930,000 SNPs then you would expect to have 7e-5 × 930,000 = 66 genotypes that are mistyped (assuming a Poisson distribution the 95% uncertainty interval is 51 – 83).
However, not all of these SNPs are “interesting”. In their marketing material 23andMe state they report on 240+ diseases and traits. Some of these involve looking at multiple SNPs, but tos simplify if we assume each disease looks at only 1 SNP, then the probability of having an error in one of the reported SNPs is 7e-5 × (240/930,000) = 1.8e-8.
With this only dpois(q=0, lambda=930000 * 1.8e-8) = 1.7% of customers will have a typing error that gives them a wrong result on 1 or more reports. With the 2829 Interpretome loci, this creeps up to 18% (again that is probability of having 1+ errors in 2829 reports).
If no calls are also considered, then the probability of 1 or more missing or wrong results per customers does approach the 1/3 that Lior calculated.
If the miscall rate is as above and the nocall rate is 1/1000, then the probability of either is 1.8e-8 × (240/930,000) + 1/1000 × (240/930000) – 2 × (1.8e-8 × (240/930000) × 1/1000 × (240/930000)) = 2.6e-7
With this rate then pois(q=0, lambda=930,000 * 2.6e-7) = 21% of customers will have at least 1 wrong or missing report. This is a far cry from “all wrong.”
December 1, 2013 at 5:24 am
Mark James Adams (@mja)
Correction: the function calls should both read
ppois(q=0, lambda=, lower.tail=F)
December 1, 2013 at 10:30 am
Lior Pachter
My assertion that all genotypes are wrong is simply based on the assumption that the number of errors is Poisson. Assuming 100 errors on average, the probability of no errors is approximately e^(-100), which I rounded up to 10^(-40). Even if one assumes less errors on average it is still the case that no individual has ever been perfectly genotyped. To be clear, I claimed “wrong genotype”, not “wrong report”.
Your calculation about the probability of a wrong or missing report is what I was alluding to in my comment that about 1/3 of tested individuals have an error at an “interesting” SNP. By such a SNP I meant one appearing in a GWAS study, and I took the number 3000 from the max of the Interactome report and the NIH GWAS catalog. I took the max since the number of SNPs is growing quickly; in fact I imagine it will be much higher in the future given the number of phenotypes for which GWAS being performed. If you believe the ENCODEies, then 80% of the sites in the genome are functional! (but I don’t believe them). What is probably true is that there are likely thousands of actionable SNPs in the genome (yet to be identified).
Your calculation breaking down results by report rather than SNP is very nice, although according to Nick Eriksson (see comment above) probably producing an overestimate of current error since he points out that the SNPs related to the report have much lower than the average error rate. The whole point of my post is that as a customer, I’d like to know what the error rates really are and I’d like to see the types of calculations you and I have been performing done by the company, and not on the back of an envelope. 23andme is probably doing them as the people working there are excellent, but one shouldn’t have to take it on faith, and it stands to reason that 23andme is not going to be the only company in the genotype interpretation space.
Returning to the claim that “all genotypes are wrong”. Some friends have asked me whether this is a relevant comment to make; after all, there is no perfect medical diagnostic test. But the point with typical tests is that maybe 99% of people get a correct answer, and 1% a wrong answer. Here every single person gets an incorrect answer overall, although of course only a small part of it is incorrect. Its sort of apples and oranges. The question is therefore how small is that small part, given the extent of testing, and your calculation makes it clear that there are different ways to estimate that. More importantly, its not at all clear that its less than 1% (or whatever threshold deemed suitable).
A final point to make is that array based genotyping is likely to be replaced by sequencing in the near future, and that introduces a whole other host of issues.
Thanks again for your comment.
December 1, 2013 at 11:07 am
Mark James Adams (@mja)
Thanks for the clarification on the meaning of ‘genotype’ in the post title. I concur that 23andMe should do a better job of communicating the technical details of the error rates of genotyping and uncertainty in their meaning for reports (such as CIs on the odds ratios) rather than us having to infer them from a few blog posts about results from MZ twins or people who had themselves genotyped twice.
December 1, 2013 at 10:51 am
Obi Griffith
When Lior titled his post “23andme genotypes are *all wrong*” I presume he was using the definition of genotype that reads something like: “the genetic constitution of an individual”. In this case it would be the complete combination of alleles at all 930,000 positions genotyped. Therefore, if even one SNP is miscalled then the genotype is technically *wrong*. Given the error rate of the technology it does follow that all of 23andme genotypes at the individual level will then be technically wrong. However, the term genotype is often defined and generally interpreted as “the genetic makeup of a cell, an organism, or an individual *usually with reference to a specific characteristic under consideration*”. When I see the term genotype I typically think of the latter sense and when I first read the title I thought – that can’t be right. Almost all of 23andme’s genotypes, as they refer to each specific characteristic, will be correct. But, reading through the post you can see how Lior meant it. So, the title while technically correct is a little bit dramatic. But hey, its a blog, and a punchy title can attract more interest. It seems to have worked in this case. The genotype at the individual level being wrong is a little bit beside the point. The real issue that at least some number of calls will be incorrect and some percentage of this time this could effect interesting SNPs. Hopefully Lior will comment on how his back of the envelope calculations differed from yours. If he also considered nocalls then his estimate of 33% isn’t that different from yours of 21%. He probably just used a different estimate of the error rate or number of interesting SNPs. Even if it is 33% of individuals getting a miscall at at least one “interesting” SNP, most of these so-called “interesting” SNPs aren’t really that interesting in a medical sense. For example, my most interesting result (sorted by risk) is that I have a 1.28x increased risk of diabetes. Without taking into account my diet, lifestyle, etc this is pretty abstract. And I would never make any medical decision about it without first contacting my doctor. I suspect that the real world chance that a 23andme result causes someone to do or not do something medically relevant is very low. But, nevertheless as these technologies improve and do become more relevant we need to think about how they are marketed and used.
December 1, 2013 at 10:23 am
Michael Eisen
I think it’s a mistake to focus on genotyping error, as the technology is improving, and this is a relatively easy to correct issue moving forward. Or to focus on error at all.
The bigger issue is not error, or the fact that these data are not being interpreted against the right background. Rather it is uncertainty – the missing information, both from the fact that we have, for most traits, an incomplete picture of the genetic basis for heritable variation in the trait, and our almost complete inability to contextualize genetics in the context of other data (environment, other risk factors, etc…) meaning that there is a very non-trivial probability that the reported risk for someone using perfect genotypes and the right background population could be wrong in magnitude and sign.
This, to me, is the real challenge. How do you convey the real range of risks to people? It’s not simply a matter of odds ratios, because they don’t capture the uncertainty of the estimate. And even this is impossible to do accurately because we don’t know what the underlying distribution of genetically attributable risk is in the population, and we never know. If I were the FDA and were interested in really protecting consumers, I would focus on working with the genetics community to develop a kind of nutrition label for genetic tests that included various forms of useful contextualizing information.
December 1, 2013 at 10:49 am
Lior Pachter
I think my post makes it clear that I’m not saying error is the only issue to worry about. I’m saying that even error is an issue to worry about (due to multiple testing).
However you make good points. I agree with all the issues you raise. One thing I’ve learned from my post and the response to it, is that there are so many issues in performing and interpreting genetic testing (on a genome-wide scale) that its inconceivable that naïve consumers have a clue, even assuming best intentions on the part of the scientists doing the basic research, DTC companies, and regulatory agencies such as the FDA.
December 2, 2013 at 7:05 pm
jjj
Hi Lior, I agree with your conclusion that the probability of a false positive is high conditioned on observing the presence of a risk-associated SNP. However I don’t think that multiple comparisons is the best explanation.
Even if only a _single_ SNP was reported, the posterior probability of an observed disease variant being a sequencing error would still be much higher than the error rate due to the base rate of risk-associated SNPs occurring is low. The reason is the low prior probability of a disease SNP being observed (I’m using prior probability in the sense of applying bayes rule in a frequentist setting where the prior corresponds to a base rate). If, hypothetically, the base rate of a risk-associated SNP was artificially high (say .5), then the posterior probability of an observed disease SNP being a sequencing error would be low.
Multiple comparisons does come into play depending on how one poses the statistical question here, but I think it doesn’t get at the crux of the inferential problem here.
In general, I’m not a fan of how multiple comparison adjustments are often used to “fix” base rate issues in GWAS inferences (my preference is more in line with M Stephens’s 2009 Nat Genetics review), but this is a bit off topic.
Anyway thanks for the insightful points and discussion.
December 2, 2013 at 6:05 am
Manuel Corpas (@manuelcorpas)
A bit late to the party but perhaps you might be interested in the article where we calculate the error rate from 23andMe chips using the genotypes of a whole family.
23andMe reports a 98% or greater call rate11, meaning that the chip can provide accurate data for more than 98% of those variants in any particular person. When an allele variant present in heterozygous state is “undercalled” (not observed), the locus may be reported as being homozygous for the other variant, leading to missed heterozygosity. Such sites may significantly impact the disease risks predicted for the individual. Under the simplifying assumption of uniform undercall probability, we estimated the number of heterozygous sites mistakenly reported as homozygous. This means that for the II-1 individual (son), 1 in every 400 sites is mistakenly called. For I-2 (father), 1 in every 200.
We also analyzed Mendelian Inheritance Errors (MIEs). If at one site a reported genotype is ‘CC’ but the genotypes for both parents is ‘TT’, one possible explanation is that one of the parents is actually heterozygous ‘CT’ but was undercalled as ‘CC’, and likewise the son is heterozygous ‘CT’ but was undercalled ‘TT’. Given 5 people in this analysis, there are 10 possible pairwise relations. Four of these represent direct parent/offspring relations, for which discrepancies can be counted as MIEs (Table 3). – See more at: http://f1000research.com/articles/1-3/
Cheers
Manny
December 5, 2013 at 11:20 pm
Konrad Karczewski
Lior,
I’m a bit late to the issue, but as the creator of Interpretome, I thought I should comment. I should first point out that that is the actual intention of Interpretome is to educate individuals with their genetic data about risk and how 23andMe arrives at their calculations. It is a teaching tool that is particularly more powerful with our book on the topic (Exploring Personal Genomics), or as part as a course such as the one we taught at Stanford. The 2 SNPs you noted from the disease tab are indeed simply from the GWAS catalog, an intersection anyone can do given time, energy, and desire. We do not provide any unified analysis for these variants and we are adamant that these are not really informative, particularly interpretable, or approved by the FDA.
Overall, I agree that the issues personal genomics is now facing are very real. Regarding error, it is true that the probability of an error in an “interesting” variant in any given individual increases with more and more interesting variants. However, in 23andMe’s case, with primarily variants of small effect for common diseases, I do really think it is the risk estimates that should be the focus of everyone’s time and attention, more than the chip error rate. Yes, for 3K variants and an error rate of 1/10K, there’s a reasonable chance that one of those variants will be miscalled. But given the distribution of mostly small effect sizes for these GWAS hits, the odds that it will make a quantifiable difference in your risk report is vanishingly small: for a variant of OR = 1.1, this error will be drowned out by the 15 other variants that have OR ~= 1.1; it would indeed be a problem if you happen to hit one of the few variants in the genome that are OR ~2, but because there are only a handful of those, the odds of that happening are small. Indeed, for large effect size variants as in rare disease, it’s a huge problem that we’ve been struggling with for years (analogies of needles in stacks of needles comes to mind), but this is only a problem if 23andMe gets into the WGS game.
I particularly agree with Michael Eisen’s comment, but would add that focusing on error rates (I realize you wrote about other things too, but the fairly sensational title is the first thing people read and focus on) only serves to distract from the bigger issues at hand. Presentation of information, and related to this, education of consumers is very important. 23andMe does a reasonable, but obviously not perfect, job of presenting their results, but I agree the uncertainty measures should be better explained. The concept of risk is of course tricky and it is difficult to highlight the importance of genetics without overstating it: I’m sure 23andMe struggles with the best way to get this across.
December 6, 2013 at 12:10 am
Lior Pachter
Thanks for your comments. Overall I think you make good points but first: if your concern is presentation of information and education of consumers, then shouldn’t Interpretome remove its Warfarin dosage calculator? The site itself calls it a “guide” and even though there are disclaimers and advice to talk to a doctor before popping pills, isn’t it irresponsible to present such results to (mostly) naïve users? Especially since recent papers suggest that previously published associations may not even be correct (see comment by Dan Reghecampf above).
I’d like to defend my choice of title. I realize it is provocative. But the facts are that
(a) it is technically correct. In fact, by the standard definition of genotype it is true that currently array and sequencing technologies cannot call a genotype completely accurately. In fact, for this reason it is impossible to know how many SNP differences there are between identical twins due to somatic mutation (there are estimates, but no precise answers). We have to acknowledge that genotyping and sequencing technology is still far from perfect.
b) I raised the multiple testing issue as it pertains to error partly to make the point that even something as simple as error, that people think is under control, is non-trivial when it comes to multiple testing. All the arguments you made about the numbers with 23andme’s current reports might be true, but we don’t really know in the absence of a calculation that is open, transparent and clear. I’m sure 23andme worries a lot about error and has mitigated it, and I believe them and take them at their word (see Nick Eriksson post above) that its not a problem, but I’d like to know for sure. You say “the odds of that happening are small”. How small? If 1 million customers are genotyped, how many get an incorrect SNP call that has major medical implications? Maybe literally zero, but the information DTC genetics companies provide can change lives. Its not a game, and the public has to have trust based on hard evidence, not faith based on myth.
c) I realize that it seems petty of me to focus on errors in genotyping, when SNP calls are 99.99.. % accurate and yet GWAS is burdened with dozens of issues that make interpretation of odds ratios a huge problem. But sometimes it pays off to think about what look like the petty issues. For example, I’ve written a paper about systematic error in sequencing that emerged from exactly such “petty” considerations: http://www.biomedcentral.com/1471-2105/12/451/
Illumina brushed it off (and proceeded to argue that it was fixed with later chemistry) but please judge for yourself by looking at the citations. Similarly, I think the issue I raised about error is important. I never said anywhere in my post it was the only, or most important issue (please see my boldface). But I think it is an important issue, and I stand by it.
I very much agree with your statement that “The concept of risk is of course tricky and it is difficult to highlight the importance of genetics without overstating it”. For this reason I appreciate the goals and implementation of Interpretome (with the exception of the Warfarin calculator), and also the role of 23andme. Perhaps it was not clear in my post, but I would very much like to see 23andme succeed. I just want to see it happen in a way that ensures responsible personal genomics in the future, when many more SNPs, technologies and companies become involved.
Thanks again for taking the time to comment, and for developing Interpretome.
December 21, 2013 at 3:53 am
Edouard Debonneuil
To make it simple, it seems we end up with:
— no 23andme genotype is perfect but that is not a problem for the non specialist users (such errors do not materially affect 23andme’s indications of risks + the error rate is goign down with time)
— the biggest issue is of course not genotyping and the uncertain conclusions we draw and present from genotyping.
— sensitivity to warfarin being a potential life-saving information for many persons, knowing **what test should be done and how to inform emergencies** about who should receive a low/medium/high dose might be VERY important (eg like the development of automated heart fibrilators).
Anyone here expert on this warfarin-sensitivity-test-and-communication question?
January 28, 2015 at 11:31 pm
deborah wilkin
bravo!
January 28, 2015 at 10:35 pm
deborah wilkin
Oh PLEASE! Regardless or error rates etc- the information 23and Me provides is amazing and wonderful. If you find a SNP in an area where you have concerns you would need to proceed to have the result validated. Simple. Only an idiot would not follow up and get further testing on “significant” SNPs. Luckily, I am not retarded….like either the editor is- or thinks we are.
January 28, 2015 at 11:28 pm
deborah wilkin
AND- their are NO issues regarding “personal genomics”- it is the consumer’s right to know available information about themselves- not yours, are any controlling bodies, right to protect us from such information- accurate or not. We are quite capable of understanding the limitations. The ppl that aren’t i.e. those that sued 23andMe for “inaccurate” information- didn’t read the disclaimer (certainly the courts threw the suit out) or were simply looking for someone to sue (or work for the FDA). Americans do not want to be saved from themselves- by you guys…. you are not near as smart or competent as you think you are- or want us to think you are- and your motivations are HIGHLY suspect…..mind your own “personal” business.
September 3, 2015 at 12:15 am
GR
I located 3 miscalls vs. a CLIA test of 20 genes. That seems pretty horrible to me to miss on some of the most important genes.
Is there any way to get a more accurate read of health genes than 23 and me?
They ignored me when I pointed it out. I’m a former engineer and health professional trying to learn about genetics, and I find the error rate troubling.